NCBI BLAST+ against the MAdLand
| Author(s) |
|
| Reviewers |
|
OverviewQuestions:
Objectives:
What is MAdLand DB?
How can we perform Blast analysis on Galaxy?
Requirements:
Load FASTA sequence into Galaxy
Perform NCBI-Blast+ analysis against MAdLandDB
Time estimation: 15 minutesSupporting Materials:Published: Jan 16, 2023Last modification: Jul 31, 2024License: Tutorial Content is licensed under Creative Commons Attribution 4.0 International License. The GTN Framework is licensed under MITpurl PURL: https://gxy.io/GTN:T00238rating Rating: 4.0 (1 recent ratings, 3 all time)version Revision: 9
MAdLandDB is a protein database comprising of a comprehensive collection of fully sequenced plant and algal genomes, with a particular emphasis on non-seed plants and streptophyte algae. Additionally, for comparative analysis, the database also includes genomes from various other organisms such as fungi, animals, the SAR group, bacteria, and archaea. The database is actively developed and maintained by the Rensing lab and released in the MAdLand setting. It employs a system of species abbreviation using a 5 letter code, which is constructed using the first three letters of the genus and the first two letters of the species name, for example, CHABR for Chara braunii. Furthermore, the database provides gene identification through the addition of gene ID’s and supplementary information such as the encoding source of the gene, whether it is plastome encoded (pt) or transcriptome-based (tr) in cases when a genome is not yet available. The key advantage of this database is its non-redundant nature, and the fact that all sequences are predominantly from genome projects, thereby increasing their reliability.
AgendaIn this tutorial, we will deal with:
Get data
Hands On: Data Upload
Create a new history for this tutorial and give it a proper name
To create a new history simply click the new-history icon at the top of the history panel:
- Click on galaxy-pencil (Edit) next to the history name (which by default is “Unnamed history”)
- Type the new name
- Click on Save
- To cancel renaming, click the galaxy-undo “Cancel” button
If you do not have the galaxy-pencil (Edit) next to the history name (which can be the case if you are using an older version of Galaxy) do the following:
- Click on Unnamed history (or the current name of the history) (Click to rename history) at the top of your history panel
- Type the new name
- Press Enter
Import the file
query.faafrom Zenodohttps://zenodo.org/records/7524427/files/query.faa
- Copy the link location
Click galaxy-upload Upload at the top of the activity panel
- Select galaxy-wf-edit Paste/Fetch Data
Paste the link(s) into the text field
Press Start
- Close the window
We just imported a FASTA file into Galaxy. Now, the next would be to perfrom the BLAST analysis against MAdLandDB.
Perform NCBI Blast+ on Galaxy
Since MAdLandDB is the collection of protein sequences, You can perform BLASTp ( Galaxy version 2.10.1+galaxy2) and BLASTx ( Galaxy version 2.10.1+galaxy2) tools.
Hands On: Similarity search against MAdLand Database
- BLASTp ( Galaxy version 2.10.1+galaxy2) OR BLASTx ( Galaxy version 2.10.1+galaxy2) with the following parameters:
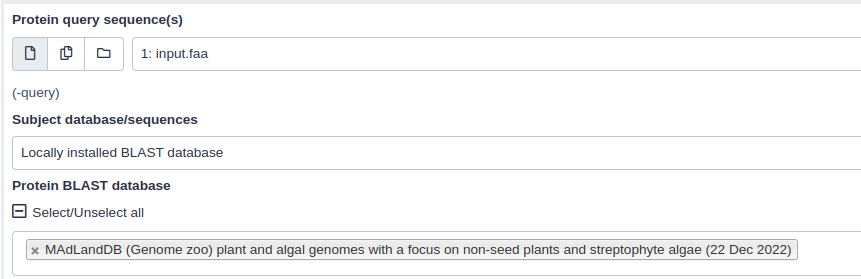
- “Protein query sequence(s)”:
Amino acid input sequence(In case of BLASTp) OR- “Translated nucleotide query sequence(s)”:
Translated nucleotide input sequence(In case of BLASTx)- “Subject database/sequences”:
Locally installed BLAST database- “Protein BLAST database”:
MadLandDB (Genome zoo) plant and algal genomes with a focus on non-seed plants and streptophyte algae (22 Dec 2022)- “Set expectation value cutoff”:
0.001- “Output format”:
- In “Output Options”:
Tabular (extended 25 columns)
Blast output
tool The BLAST output will be in tabular format (you can select the desired output format from the drop down menu) and include the following fields :
| Column | NCBI name | Description |
|---|---|---|
| 1 | qseqid | Query Seq-id (ID of your sequence) |
| 2 | sseqid | Subject Seq-id (ID of the database hit) |
| 3 | pident | Percentage of identical matches |
| 4 | length | Alignment length |
| 5 | mismatch | Number of mismatches |
| 6 | gapopen | Number of gap openings |
| 7 | qstart | Start of alignment in query |
| 8 | qend | End of alignment in query |
| 9 | sstart | Start of alignment in subject (database hit) |
| 10 | send | End of alignment in subject (database hit) |
| 11 | evalue | Expectation value (E-value) |
| 12 | bitscore | Bit score |
The fields are separated by tabs, and each row represents a single hit. For more details for BLAST analysis and output, we recommand you to follow the Similarity-searches-blast tutorial.
See Cock et al. 2015 and Cock et al. 2013
More Similarity Search Tools on Galaxy
- Diamond: Diamond ( Galaxy version 2.0.15+galaxy0) is a high-throughput program for alignment of large-scale data sets. It aligns sequences to the reference database using a compressed version of the reference sequences called a “database diamond” which is faster to read and can save computational time (~20,000 times the speed of Blastx, with high sensitivity).
See Buchfink et al. 2014 for more discussion.