Creating content in Markdown
| Author(s) |
|
| Editor(s) |
|
OverviewQuestions:
Objectives:
How to write a tutorial with hands-on?
What are the different boxes?
How can I add a caption to an image?
Create hands-on
Use the different boxes
Time estimation: 15 minutesSupporting Materials:Published: Jun 25, 2017Last modification: Jan 27, 2024License: Tutorial Content is licensed under Creative Commons Attribution 4.0 International License. The GTN Framework is licensed under MITpurl PURL: https://gxy.io/GTN:T00058version Revision: 123
Once we have set up the infrastructure, we are ready to write the tutorial.
AgendaIn this tutorial, we will cover:
The tutorial’s content should be placed in the file tutorial.md. Its syntax and structure are simple, and will have the following structure:
---
layout: tutorial_hands_on
title: Title of the tutorial
zenodo_link: ''
questions:
- Which biological questions are addressed by the tutorial?
- Which bioinformatics techniques are important to know for this type of data?
objectives:
- The learning objectives are the goals of the tutorial
- They will be informed by your audience and will communicate to them and to yourself
what you should focus on during the course
- They are single sentences describing what a learner should be able to do once they
have done completed tutorial
- You can use Bloom's Taxonomy to write effective learning objectives
time_estimation: ''
key_points:
- The take-home messages
- They will appear at the end of the tutorial
contributors:
- contributor1
- contributor2
---
introduction text
> <agenda-title></agenda-title>
>
> In this tutorial we will deal with:
>
> 1. TOC
> {:toc}
# Section 1
blabla
## Subsection 1
blabla
# Section 2
blabla
## Subsection 2
blabla
# Conclusion
blabla
Metadata
The tutorial.md needs to start with some metadata at the top:
layout: tutorial_hands_on: keep the defaulttitle: title of the tutorial (it will appear on the tutorial page and the topic page)level:Introductory,IntermediateorAdvancedzenodo_link: link on Zenodo to the input data for the tutorialcontributions: eveybody who has contributed to this tutorial (usernames must match those inCONTRIBUTORS.yamlfile)
Hands-on: Fill the basic metadata
Update the tutorial information in the header section of your tutorial:
title: "Similarity search with BLAST"(Optional) Add the Zenodo link (if created)
This information is used to display the data from the topic and tutorial page. They are also used to check which information is missing for the tutorials.
We also define metadata related to the pedagogical content of the tutorial, which will appear at the top (“Overview” box) and bottom of the online tutorial:
A dictionary/map
Free Textlayout
This must be set to
tutorial_hands_onPossible Values:
tutorial_hands_onExample(s)
layout: "tutorial_hands_on"Free Texttitle
Title of the tutorial (it will appear on the tutorial page and the topic page)
Example(s)
title: Clustering in Machine Learningtitle: Breve introducción a Galaxy - en españoltitle: Pangeo ecosystem 101 for everyone - Introduction to Xarray Galaxy ToolsA dictionary/mapcontributions
List of tutorial contributors. Here we break them down into several broad categories to help contributors identify how they contributed to a specific tutorial.
Example(s)
contributions: authorship: - shiltemann - bebatut editing: - hexylena - bebatut - natefoo testing: - bebatut infrastructure: - natefoo translation: - shiltemann funding: - gallantriesList of Items
These entities wrote the bulk of the training material, they may have done the analysis, built the workflow, and wrote the text themselves.
List of Items
These entities edited the text, either for spelling and grammar, flow, GTN-fit, or other similar editing categories
List of Items
These entities provided funding support for the development of this resource
List of Items
These entities managed and provided infrastructure to the GTN or for training purposes
List of Items
These entities tested the tutorial to ensure it works correctly for students, or reported issues with the tutorial.
List of Items
These entities did translation and localisation work on this resource
List of Items
These entities contributed UX or Design improvements to this tutorial or the GTN as a whole
Free Texttime_estimation
An estimation of the time needed to complete the hands-on.
Example(s)
time_estimation: 10Mtime_estimation: 1H30MRequired Pattern: Must match the following regular expression
/^(?:([0-9]*)[Hh])*(?:([0-9]*)[Mm])*(?:([0-9.]*)[Ss])*$/A dictionary/map
A dictionary of abbreviations and their expansions.
Example(s)
abbreviations: SQL: Structured Query Language API: Application Programming InterfaceFree Text
The expansion of the abbreviated term.
List of Items
List of tutorial contributors. Please use
contributionsinstead as it provides more detailed accounting of tutorial history.Example(s)
contributors: - hexylena - shiltemannFree Text
A copyright attribution string, as required by some licenses.
Example(s)
copyright: © Copyright 2021-2023 University of Technology Sydney, The University of Manchester UK and RO-Crate contributorsBoolean
trueto hide your tutorial from the topic page (optional). This is useful if you need a tutorial for a workshop, but have not finished making it up to GTN standards.List of Items
A dictionary/map
Any additional variables you want to set on the page
List of Items
list of resources that the reader of the material could follow at the end of the tutorial
Example(s)
follow_up_training: - type: internal topic_name: statistics tutorials: - age-prediction-with-ml - type: external title: The Unix Shell link: "http://swcarpentry.github.io/shell-novice/" - type: none title: "A VM with at least 2 vCPUs and 4 GB RAM, preferably running Ubuntu 18.04 - 20.04."A dictionary/mapSequence Value (List of items)
Free Texttype
the type of link
Possible Values:
internalexternalnoneExample(s)
type: "internal"type: "external"type: "none"Free Text
URL of the external resource
Free Text
Title of the external resource
Free Text
[Internal Only] The name of the topic
List of Items
[Internal Only] List of required tutorials inside that topic
Decimal Number
Currently unused.
Free Text
Link to a gitter channel that is more relevant for the tutorial than the default. E.g. a single cell tutorial could use
Galaxy-Training-Network/galaxy-single-cellto link to their specific chat room.Example(s)
gitter: Galaxy-Training-Network/galaxy-single-cellgitter: galaxy-genome-annotation/LobbyFree Text
This must be set to
externalto link to an external tutorialPossible Values:
externalExample(s)
hands_on: "external"Free Text
link to the external tutorial
Example(s)
hands_on_url: https://docs.qiime2.org/jupyterbooks/cancer-microbiome-intervention-tutorial/index.html#List of Items
List of take-home messages. This information will appear at the end of the tutorial. These should really be a key point, something that should stick in their mind; what you want them to take home from the tutorial.
Example(s)
key_points: - Pangeo ecosystem enables big data analysis in geosciences - The MiModD suite of tools bundles most of the functionality required to perform mapping-by-sequencing analyses with Galaxy - It can drastically simplify management of large numbers of VMsFree Text
The document language.
Possible Values:
esenfrExample(s)
lang: "es"lang: "en"lang: "fr"Free Text
Here give a feeling of what level the material is at.
Possible Values:
IntroductoryIntermediateAdvancedExample(s)
level: "Introductory"level: "Intermediate"level: "Advanced"Free Text
An SPDX identifier for the alternative license that is used for that particular material. This is only relevant for materials which have been imported from an external source and were originally licensed under another license. For new materials we strongly encourage contributors to not use this key as materials are CC-BY, by default.
Example(s)
license: MITlicense: Apache-2.0A dictionary/map
Example(s)
notebook: language: python pyolite: true notebook: language: python snippet: topics/climate/tutorials/pangeo-notebook/preamble.mdFree Textlanguage
Possible Values:
pythonbashrsqlExample(s)
language: "python"language: "bash"language: "r"language: "sql"List of Items
A list of packages that must be installed before running this tutorial. This value is not currently used, but might be in the future.
Example(s)
packages: - tidyverseBoolean
The GTN has support for JupyterLite and the Pyodide kernel which runs Python in the browser via webassembly/javascript. This comes with some restrictions:
- Python only
- No filesystem access (so no
wgetprep steps)- Little to no cell magic
However, it means we can run a lot of our Python training directly in the GTN! And in the future, hopefully, we will be able to embed individual cells of the notebook directly in the Python training, so the user doesn’t even need to switch pages.
Enabling this field will enable pyolite links for this notebook.
Free Text
If you have an alternative preamble for your notebook that students should know before following (e.g. they must load X datasets in their history), it can be listed here.
This text will be shown in the GTN tutorial, but it will not be included in the notebook, giving you a bit better control over mixing setup content which is relevant for Galaxy, with notebook content that can be easy to run for students.
Example(s)
snippet: topics/climate/tutorials/pangeo-notebook/preamble.mdList of Items
list of learning objectives for the tutorial
A learning objective is a single sentence describing what a learner will be able to do once they have done the tutorial. Generally it is best to follow a 2C or 3C learning objective such as:
- Compute (Skill)
- multiple whole genome assemblies (Objective)
- in such a way to develop big data processing skills (Result)
Example(s)
objectives: - Understand the basic concepts behind phylogenetic trees, as applied to *Mycobacterium tuberculosis* - Explore Biodiversity data with taxonomic, temporal and geographical informations - Generate a DotPlot emulating the original paper using a different analysis toolFree Text
A custom image to show on the link preview in external applications (e.g. when the URL is pasted into Twitter)
Example(s)
og_image: /assets/images/gat.pngRequired Pattern: Must match the following regular expression
/^\/.*/Integer Number
This field allows ordering tutorials within the tutorial list. Tutorials with lower numbered priority come before tutorials with higher numbers.
Example(s)
List of Items
list of questions that will be addressed in the tutorial
Example(s)
questions: - How does Genome assembly work? - How do I change Galaxy configs? - How to detect and quantify differentially abundant proteins in a HEK-Ecoli Benchmark DIA datatset? - What kinds of data do programs store?List of Items
If a tutorial is renamed to a new location, use this field to list prior locations from which this tutorial was accessible.
Example(s)
redirect_from: - /topics/sequence-analysis/tutorials/de-novo-rad-seq/tutorialList of Items
List of resources that the reader of the material should be familiar with before starting this training. The structure is identical to
follow_up_training.Example(s)
requirements: - type: internal topic_name: statistics tutorials: - age-prediction-with-ml - type: external title: The Unix Shell link: "http://swcarpentry.github.io/shell-novice/" - type: none title: "A VM with at least 2 vCPUs and 4 GB RAM, preferably running Ubuntu 18.04 - 20.04."A dictionary/mapSequence Value (List of items)
Free Texttype
the type of link
Possible Values:
internalexternalnoneExample(s)
type: "internal"type: "external"type: "none"Free Text
URL of the external resource
Free Text
Title of the external resource
Free Text
[Internal Only] The name of the topic
List of Items
[Internal Only] List of required tutorials inside that topic
Free Text
if the topic has multiple subtopics defined, you can assign your tutorial to one of those subtopics here. Without this, the tutorial will appear in the “Other tutorials” section on the topic page.
Example(s)
subtopic: single-cellList of Items
A free form list of tags that are relevant for your tutorial.
Example(s)
tags: - covid-19 - git-gatList of Items
If alternative translations of a material are available, then use this key to indicate which languages have been manually translated.
Example(s)
translations: - enA dictionary/map
For materials which are automatically converted into videos via the available mechanisms, this field declares which voice should be used. If this field is not declared, a random voice will be chosen from a list of the best available voices from AWS Polly.
Example(s)
voice: id: Lupe lang: es-US neural: trueFree Textid
Free Textlang
Booleanneural
Free Text
link on Zenodo to the input data for the tutorial
Example(s)
zenodo_link: https://zenodo.org/record/3706539Required Pattern: Must match the following regular expression
/(^$|^https://zenodo.org/records?/[0-9]+/?$|^https://doi.org/10.5281/zenodo.[0-9]+/?$)/
For this category of metadata, we have taken inspiration from what Software Carpentry has done and particularly what they described in their Instructor training.
Hands-on: Fill out the pedagogical metadata
- Define 2 questions that will be addressed during the tutorial and add them to the metadata
- Define 2 learning objectives for the tutorial and add them to the metadata
Comment: When filling the pedagogical metadataWe recommend that you fill out the questions and the learning objectives before starting writing the tutorial content. You can still refine them afterwards, but it will help to guide you in developing your tutorial, and gives you some time to think beforehand on what topics are worth being covered.
For the take-home messages, it is easier to define them once the tutorial is written and you identified the issues.
Listing contributors
All tutorials and slides must give credit to all contributors. This can be any type of contribution, adding them in GitHub, creating images for it, etc.
-
Make sure all contributors are listed in the
CONTRIBUTORS.yamlfile. Each contributor is defined in this file like:contributor-username: # GitHub username (if the contributor has one) name: Full Name # mandatory joined: 2020-06 # mandatory email: saskia.hiltemann@gmail.com # optional twitter: shiltemann # optional linkedin: shiltemann # optional gitter: shiltemann # optional orcid: 0000-0003-3803-468X # optional bio: Researcher at EMC # optional -
Add all contributors to the metadata of the tutorial or slide deck:
contributors: - contributor-username - shiltemann - hexylenaMake sure these names match the usernames you used in the
CONTRIBUTORS.yamlfile. -
Optional: Specifying types of contributions. If you want to give more detailed credit for conributions, you can do the following (instead of step 2 above)
contributions: authorship: - shiltemann editing: - bebatut - hexylena funding: - carpentries testing: - userX ux: - userY infrastructure: - userZTo define a funding body in the
CONTRIBUTORS.yamlthere are a few extra fields available:gallantries: name: Gallantries Project joined: 2020-09 avatar: "https://www.erasmusplus.nl/assets/images/logo.png" github: false funder: true funding_id: 2020-1-NL01-KA203-064717 funding_statement: | This project ([`2020-1-NL01-KA203-064717`](https://erasmus-plus.ec.europa.eu/projects/search/details/2020-1-NL01-KA203-064717)) is funded with the support of the Erasmus+ programme of the European Union. Their funding has supported a large number of tutorials within the GTN across a wide array of topics. <a href="https://gallantries.github.io/assets/images/logosbeneficaireserasmusright_en.jpg" rel="noopener noreferrer"><img src="https://gallantries.github.io/assets/images/logosbeneficaireserasmusright_en.jpg" alt="eu flag with the text: with the support of the erasmus programme of the european union. " loading="lazy"></a>Funding bodies will be credited at the bottom of the tutorial with the appropriate funding statement, and will get a page in the hall of fame listing all tutorials that list them as a funder.
For an example of how this all looks, see the R basics tutorial (top and bottom of the tutorial).
A dictionary/mapContributions Schema
List of tutorial contributors. Here we break them down into several broad categories to help contributors identify how they contributed to a specific tutorial.
Example(s)
contributions: authorship: - shiltemann - bebatut editing: - hexylena - bebatut - natefoo testing: - bebatut infrastructure: - natefoo translation: - shiltemann funding: - gallantriesList of Items
These entities wrote the bulk of the training material, they may have done the analysis, built the workflow, and wrote the text themselves.
List of Items
These entities edited the text, either for spelling and grammar, flow, GTN-fit, or other similar editing categories
List of Items
These entities provided funding support for the development of this resource
List of Items
These entities managed and provided infrastructure to the GTN or for training purposes
List of Items
These entities tested the tutorial to ensure it works correctly for students, or reported issues with the tutorial.
List of Items
These entities did translation and localisation work on this resource
List of Items
These entities contributed UX or Design improvements to this tutorial or the GTN as a whole
A dictionary/map
Example(s)
hexylena: name: Helena twitter: hexylena bio: I wrote this documentation! I do super cool things.A dictionary/map
This ideally is your GitHub handle. If you do not have, or do not wish to provide a GitHub username, you may make up another identifier here, but then you must set
github: falseas described below.Free Textname
Your preferred name. If you prefer an alias or another name, this is welcome, it does not need to be your legal name.
Example(s)
name: 张三name: Alicename: Jane Doename: Madame Tout-le-Mondename: Γιάννης ΠαπαδόπουλοςFree Textjoined
The year and month in which you joined
Example(s)
joined: 2020-01Required Pattern: Must match the following regular expression
/[0-9]{4,}-[0-9]{2}/List of Items
A set of organisations or grants your are affiliated with
Free Text
Free Text
A short biography of yourself, if you wish to add additional details or context.
Example(s)
bio: Research at the [South African National Bioinformatics Institute](https://www.sanbi.ac.za/)Free Text
Your bluesky handle
Example(s)
bluesky: gtn.bsky.socialRequired Pattern: Must match the following regular expression
/[0-9a-zA-Z.]+/Boolean
Are you open to being contacted for hosting training, or being involved with local training events? Instructors might use this to contact you, if they are looking for regional trainers who can help with their event.
Free Text
The 2 letter code identifying the ELIXIR node to which you are a member or are associated with. If you are from norway, you will need to quote your value,
"no", unlike everyone else, due to the Norway Problem with YAMLPossible Values:
aubechczdedkeeesfifrgrhuieilitlunlnoptsesiukExample(s)
elixir_node: "au"elixir_node: "be"elixir_node: "ch"elixir_node: "cz"elixir_node: "de"elixir_node: "dk"elixir_node: "ee"elixir_node: "es"elixir_node: "fi"elixir_node: "fr"elixir_node: "gr"elixir_node: "hu"elixir_node: "ie"elixir_node: "il"elixir_node: "it"elixir_node: "lu"elixir_node: "nl"elixir_node: "no"elixir_node: "pt"elixir_node: "se"elixir_node: "si"elixir_node: "uk"Free Text
Your email address, if you wish to provide it.
Example(s)
email: jane.doe@gmail.comRequired Pattern: Must match the following regular expression
/@/Free Text
The URL to your fediverse profile
Example(s)
fediverse: http://genomic.social/@abretaudRequired Pattern: Must match the following regular expression
/^https:\/\/[0-9a-zA-Z.]+/@?[0-9a-zA-Z.]+$/Free Text
The flavor of the fediverse server (used in our webfinger endpoint.)
Possible Values:
mastodonakkomaExample(s)
fediverse_flavor: "mastodon"fediverse_flavor: "akkoma"List of Items
A set of organisations you were previously affiliated with
Boolean
If your identifier in this file is not a GitHub account (or not your account), then this must be set to true, so we do not link to that account.
Free Text
Set this to
noif you would like to be excluded from the hall of fame.Possible Values:
noExample(s)
halloffame: "no"Free Text
If this community member is deceased, you can use this field to write a small in memoriam section that will display in their Hall of Fame profile.
Free Text
Required Pattern: Must match the following regular expression
/[0-9a-zA-Z]+/A dictionary/map
Free Text
The country you work in, ISO-3166-2 code.
Example(s)
country: NLcountry: DEDecimal Number
Latitude in decimal numbers
Example(s)
lat: 4.12Decimal Number
Longitude in decimal numbers
Example(s)
lon: 55.0Free Text
Preferred contact method
Free Text
Your matrix identifier and home server
Example(s)
matrix: hexylena:matrix.orgRequired Pattern: Must match the following regular expression
/[0-9a-zA-Z]+:.*/Free Text
Your identifier at orcid.org
Example(s)
orcid: 0000-0001-9760-8992Required Pattern: Must match the following regular expression
/[0-9A-Z]{4}-[0-9A-Z]{4}-[0-9A-Z]{4}-[0-9A-Z]{4}/Free Text
Your twitter handle, without the
@Example(s)
twitter: gxytrainingRequired Pattern: Must match the following regular expression
/[0-9a-zA-Z]+/
Content
The tutorial’s content is written directly after the section of metadata. This is written in Markdown, a simple markup language.
Comment: MarkdownCheck this cheatsheet to learn more how to use Markdown.
The Markdown content is then transformed into a user friendly webpage through a templating system. With this approach, there is no need to add the name of every tutorial each time, since they are automatically added based on the tutorial’s metadata.
To help developing the tutorial, we recommend to create a workflow of the different steps of the tutorial inside Galaxy first, and then you can create the structure of the tutorial automatically from that:
Hands-on: Create the structure of the tutorial from a workflow
- Create a small workflow with one or two steps on a running Galaxy instance
- Add the topic name as Tag and the tutorial title as Annotation/Notes to the workflow using the workflow editor.
- Get the workflow id
- Go the “Share” page of the workflow
- Copy the information after
id=in the URL of the page- Get your API key for this Galaxy instance
- Click on User –> Preferences
- Click on Manage API key
- Click on Create a new key (if none is available)
- Copy the API key
Generate the skeleton of the tutorial locally
$ planemo training_generate_from_wf \ --topic_name "my-topic" \ --tutorial_name "my-new-tutorial" \ --galaxy_url "URL to Galaxy instance in which you created the workflow" \ --galaxy_api_key "Your API key on the Galaxy instance" \ --workflow_id "ID of the workflow on the Galaxy instance" \ --zenodo_link "URL to the Zenodo record (Optional)"Comment: Using a local workflowIt is also possible to download the workflow locally (with the
.gaextension), and then run a slightly different command:$ planemo training_generate_from_wf \ --topic_name "my-topic" \ --tutorial_name "my-new-tutorial" \ -- workflow PATH/to/the/file.ga \ --zenodo_link "URL to the Zenodo record (Optional)"- Inspect the generated
tutorial.md
The generated tutorial is structured with:
- An introduction, to give an overview of the tutorial with its use cases, data, and methods
- Multiple sections representing the steps of the analysis, complete with automatically generated hands-on blocks, as practicing is a vital part of the learning process
- A conclusion to summarize what has been done in the tutorial (with a graphic)
Hands-on: Filling out the structure of the tutorial
- Fill out the “Introduction” with a general introduction of the tutorial and a small description of the dataset (goals)
- Rename/restructure the sections with several levels and more explication
- Add some theory about the tool used to introduce each section
- Add a small conclusion and relate the results to the original question
Self Study Tutorials
By following all of the guidelines in this file you can be sure that your tutorial will be optimised for self-study. The GTN framework encourages the use of snippets, and PTDK ensures tutorials have fully detailed parameters learners should configure. If you make use of snippets in appropriate places, learners can easily follow a tutorial despite different skill levels with Galaxy.
Additionally use of Question and Solution boxes can ensure that students can self-check their understanding and progress as they progress through the tutorial. Generally we recommend one or more questions (and associated correct solutions!) after every hands-on box (which might have a one or more steps to follow.) These questions will let students be sure their work is correct before they proceed too far. As such you should design your questions carefully in order to catch common and likely failure modes that learners may encounter. If a student might forget to select a less-common or deeply-nested parameter, be sure that your question following that hands on tests that properly, and if possible explain that in the solutions.
Adding images with captions
To add an image in Markdown file, we need to use the markdown syntax for this: .
We have also added a small plugin to handle captions for each image:
 Open image in new tab
Open image in new tabThe prefix “Figure 1.” is automatically added before its caption. This is done with the following Markdown syntax:

We can also cross-reference images inside our Markdown with an anchor. For example, we can link to the previous figure using [the display text](#figure-nb) (changing nb with the image’s number).
Guidelines on Alt vs Figcaption Text
While both the alt attribute and the figcaption element provide a way to describe images, the way we write for them is different.
altdescriptions should be functional;figcaptiondescriptions should be editorial or illustrative.
As an example for this image:
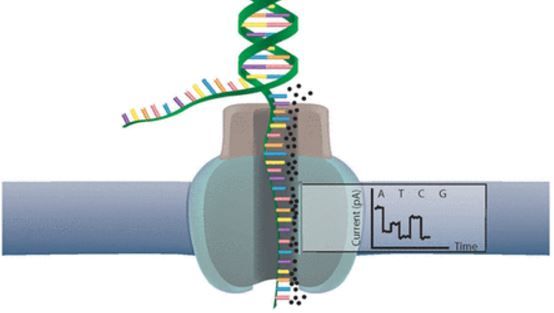 Open image in new tab
Open image in new tab
| Field | Appropriate Contents |
|---|---|
| alt text | Image of cell membrance with an embedded protein with central pore. DNA is shown splitting and entering the pore, an electrical signal comes out reading A C T or G. |
| figure caption | Using nanopore sequencing, a single molecule of DNA or RNA can be sequenced without the need for PCR amplification or chemical labeling of the sample. (Image from: Nanopore sequencing: The advantages of long reads for genome assembly) |
Writing mathematical expressions
Mathematical expressions can be written in LaTeX, and are automatically rendered with MathJax.
Surround your math expression with two $ signs on each side (like in LaTeX math blocks):
- inline expressions, e.g.
$$ 5 + 5 $$will be rendered as \(5 + 5\) -
block expressions, e.g.
\[5 + 5\]$$ 5 + 5 $$will be rendered in its own line block as
Dollar signs are therefore reserved characters for instructing the templating system to open/close LaTeX math blocks. If you want to use a $ within your expression, you will need to escape it: $$ a + 3\$ = 5\$ $$ will be rendered as: \(a + 3\$ = 5\$\)
LaTeX code that uses the pipe symbol | in inline math statements may lead to a line being recognized as a table line by the templating system.
This can be avoided by using the \vert command instead of |
Tables and Matrices
Tables can be generated using markdown by using the | symbol to indicate column dividers, and -- for table headers:
| | Obs1 | Obs2 | Obs3 |
|------ |--------------------|
| Feat1 | 0 | 1 | 2 |
| Feat2 | 1 | 2 | 3 |
| Feat3 | 2 | 3 | 4 |
When rendered, they will take the full width of the page:
| Obs1 | Obs2 | Obs3 | |
|---|---|---|---|
| Feat1 | 0 | 1 | 2 |
| Feat2 | 1 | 2 | 3 |
| Feat3 | 2 | 3 | 4 |
This does not appear to be visually appealing when representing matrices, which is why a matrix box can be used instead:
> | | Obs1 | Obs2 | Obs3 |
> | ----- |--------------------|
> | Feat1 | 0 | 1 | 2 |
> | Feat2 | 1 | 2 | 3 |
> | Feat3 | 2 | 3 | 4 |
{: .matrix}
The rendered table is then given as a minimum-width and centred matrix:
Obs1 Obs2 Obs3 Feat1 0 1 2 Feat2 1 2 3 Feat3 2 3 4
Internally linking to other training material
If you want to link to other training material within your text, please use the {% link path/to/file.ext %} tag:
[link text]( {% link topics/single-cell/tutorials/scrna-case_monocle3-trajectories/tutorial.md %} )
(Note the .md extension, and not .html, always provide the file name here, it will automatically be converted to the correct link)
If you want to link to a specific section in a tutorial using an anchor (e.g. #getting-started), place it outside of the {% link %} tag:
[link text]({% link topics/single-cell/tutorials/scrna-case_monocle3-trajectories/tutorial.md %}#section-name)
Improving the learning experience with Boxes
To improve the learning experience in our tutorial, we define some boxes to highlight content.
These boxes are defined always with the same structure:
> <type-title>Name of the box</type-title>
> text goes here!
{: .type}
where type is something like tip, so <tip-title> and {: .tip}. You must follow this structure exactly for it to be rendered correctly.
Overview box
This box at the top of each tutorial is automatically generated using the metadata we defined in the topic’s metadata file:
: Overviewquestion Questions
- Which biological questions are addressed by the tutorial?
- Which bioinformatics techniques are important to know for this type of data?
objectives Objectives
- The learning objectives are the goals of the tutorial
- They will be informed by your audience and will communicate to them and to yourself what you should focus on during the course
- They are single sentences describing what a learner should be able to do once they have completed the tutorial
- You can use Bloom’s Taxonomy to write effective learning objectives
requirements Requirements
time Time estimation: ‘1H’
Hands-on: Checking the metadata
Check that the metadata added previously are correctly filling the overview box
QuestionWhat metadata hasn’t been added to this box?
The take-home messages are not added to this box but into the last box of the tutorial
Agenda box
In most tutorials, the second box is the agenda box, placed at the end of the introduction. It shows the table of contents for the tutorial
> <agenda-title></agenda-title>
>
> In this tutorial we will deal with:
>
> 1. TOC
> {:toc}
>
{: .agenda}
There is no need to fill out the list; this will be done automatically based off of your tutorial’s section titles. First and second level headings (lines starting with # and ## in markdown) will be shown in the agenda.
If a section is not important enough to be shown in the agenda, consider making it a third-level heading (###) instead.
The agenda will be rendered as follows:
AgendaIn this tutorial we will deal with:
- TOC
Hands-on box
We find that having users walk through the tutorial, doing all of the steps is important for learning the concepts taught. We therefore emphasize this by regularly adding hands-on sections, where trainees are encouraged to do the analysis by themselves. We have designed some special boxes to make these sections easier to find.
> <hands-on-title>Spliced mapping</hands-on-title>
>
> 1. **RNA STAR** {% icon tool %}: Map your reads on the reference genome with
> - *"Single-end or paired-end reads"*: `Paired-end (as individual datasets)`
> - *"RNA-Seq FASTQ/FASTA file, forward reads"*: the generated `trimmed reads pair 1` files (multiple datasets)
> - *"RNA-Seq FASTQ/FASTA file, reverse reads"*: the generated `trimmed reads pair 2` files (multiple datasets)
> - *"Custom or built-in reference genome"*: `Use a built-in index`
> - *"Reference genome with or without an annotation"*: `use genome reference without builtin gene-model`
> - *"Select reference genome"*: `Drosophila Melanogaster (dm6)`
> - *"Gene model (gff3,gtf) file for splice junctions"*: the imported `Drosophila_melanogaster.BDGP6.87.gtf`
> - *"Length of the genomic sequence around annotated junctions"*: `36`
>
> This parameter should be length of reads - 1
>
> 2. {% tool [MultiQC](toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.8+galaxy0) %}: Aggregate the STAR logs with
> - *"Which tool was used generate logs?"*: `STAR`
> - *"Type of FastQC output?"*: `Log`
> - *"STAR log output"*: the generated `log` files (multiple datasets)
{: .hands_on}
For consistency please use:
{% icon hands_on %}icon to define that is an hands-on- Short imperative sentences to make it easy to identify the tasks
- Name of the tool in bold followed by
{% icon tool %}icon to make it easy to identify a Galaxy tool - Parameters for the tool as a sublist
This will be rendered like:
Hands-on: Spliced mapping
- RNA STAR tool: Map your reads on the reference genome with
- “Single-end or paired-end reads”:
Paired-end (as individual datasets)- “RNA-Seq FASTQ/FASTA file, forward reads”: the generated
trimmed reads pair 1files (multiple datasets)- “RNA-Seq FASTQ/FASTA file, reverse reads”: the generated
trimmed reads pair 2files (multiple datasets)- “Custom or built-in reference genome”:
Use a built-in index- “Reference genome with or without an annotation”:
use genome reference without builtin gene-model- “Select reference genome”:
Drosophila Melanogaster (dm6)- “Gene model (gff3,gtf) file for splice junctions”: the imported
Drosophila_melanogaster.BDGP6.87.gtf“Length of the genomic sequence around annotated junctions”:
36This parameter should be length of reads - 1
- MultiQC ( Galaxy version 1.8+galaxy0): Aggregate the STAR logs with
- “Which tool was used generate logs?”:
STAR- “Type of FastQC output?”:
Log- “STAR log output”: the generated
logfiles (multiple datasets)
There are also some predefined parameter icons available which can be used to indicate the type of parameter. These are optional, but may be helpful in some cases (for example to distinguish between single file inputs and collection inputs).
The available icons are:
> <hands-on-title>My Step</hands-on-title>
>
> 1. **My Tool** {% icon tool %} with the following parameters
> - {% icon param-text %} *"My text parameter"*: `my value`
> - {% icon param-file %} *"My input file"*: `my file`
> - {% icon param-files %} *"My multiple file input or collection"*: `my collection`
> - {% icon param-select %} *"My select menu"*: `my choice`
> - {% icon param-check %} *"My check box"*: `yes`
> - {% icon param-toggle %} *"My toggle button"*: `Yes`
> - {% icon param-repeat %} **My repeat parameter**
> - *"param1"*: `42`
{: .hands_on}
which, when rendered, look like:
Hands-on: My Step
- My Tool tool with the following parameters
- param-text “My text parameter”:
my value- param-file “My input file”:
my file- param-files “My multiple file input or collection”:
my collection- param-select “My select menu”:
my choice- param-check “My check box”:
yes- param-toggle “My toggle button”:
Yes- param-repeat My repeat parameter
- “param1”:
42
Questions and Solutions boxes
Questions can be added to force trainees to think about what they are currently doing, and to put things in perspective. They can also help the instructors by exposing and clarifying common scenarios, errors, or applications.
> <question-title></question-title>
>
> 1. Why are some tests filtered?
> 2. Does it improve the *p*-value distribution?
>
> > <solution-title></solution-title>
> >
> > 1. Sol for the first question
> > 2. Sol for the second question
> >
> {: .solution}
{: .question}
Which will be rendered as:
Question
- Why are some tests filtered?
- Does it improve the p-value distribution?
- Sol for the first question
- Sol for the second question
Questions should be quick to answer. You can directly ask a question and expect an answer, or you can provide some answers and create multiple choices questions (MCQs). With well chosen wrong answers, MCQs can do much more than just measure how much someone knows, such as exposing common misconceptions and mistakes.
In the box below, initially hidden, we add the correct answer and possibly any additional explanation. Self-trainees can then check the solution and its explanation.
In the context of remote trainings, where a teacher isn’t synchronously available, ensuring that you have questions throughout your materials for students to check their understanding is incredibly key.
Additionally ensuring that solutions are provided, and are correct and up-to-date (or use a snippet explaining data variability along with with ways to check the results) is mandatory. Students will then use these questions to self-check their understanding against what you expected them to learn.
Tips box
Tips boxes are really just for ‘tips’, usually hints regarding Galaxy operations that users may or may not be familiar with. If you want to provide extended discussion or links to external materials then consider the comment and detail boxes instead.
> <tip-title>Importing data via links</tip-title>
>
> * Copy the link location
> * Open the Galaxy Upload Manager
> * Select **Paste/Fetch Data**
> * Paste the link into the text field
> * Press **Start**
{: .tip}
Rendered:
- Copy the link location
- Open the Galaxy Upload Manager
- Select Paste/Fetch Data
- Paste the link into the text field
- Press Start
Comments boxes
> <comment-title></comment-title>
> - Edit the "Database/Build" to select "dm3"
> - Rename the datasets according to the samples
{: .comment}
Rendered:
Comment
- Edit the “Database/Build” to select “dm3”
- Rename the datasets according to the samples
Details box
The detail box is used to give more background explanation on the subject. By default the box is collapsed, trainees can expand it if they wish to know extra information about a topic.
> <details-title>More details on the ....</details-title>
>
> Add more details in Markdown...
>
{: .details}
Rendered:
Add more details in Markdown…
Key points box
This last box of the tutorial is automatically created with the take-home messages defined in the topic’s metadata
To render the boxes correctly, the syntax needs to be correct. If it doesn’t work, have a look at similar tutorials and get inspiration.
Warning box
> <warning-title>Danger: You can lose data!</warning-title>
> Something really bad can happen here!
{: .warning}
Rendered:
Warning: Danger: You can lose data!Something really bad can happen here!
Nested boxes
Boxes can be nested, e.g. for having tips inside a hands-on:
> <hands-on-title>Defining the topic for the tutorial</hands-on-title>
>
> 1. Search for NCBI Blast+ on the [ToolShed](https://toolshed.g2.bx.psu.edu/)
> 2. Check in which category it is
>
> > <question-title></question-title>
> >
> > In which topic will you put the tutorial?
> >
> > > <solution-title></solution-title>
> > >
> > > If we search for [NCBI Blast+ in the ToolShed](https://toolshed.g2.bx.psu.edu/view/devteam/ncbi_blast_plus/7538e2bfcd41), it is attributed to 2 categories (bottom): "Next Gen Mappers" and "Sequence Analysis".
> > > We decided to put it in "Sequence analysis" because this is the most general one for this tutorial.
> > {: .solution}
> {: .question}
{: .hands_on}
Code box
We have added code in/out boxes to help you show commands, and their effects, when running command line commands.
Normally a single column, with the boxes above one another, it will automatically split side-by-side over a given width (1200px);
> > <code-in-title>Bash</code-in-title>
> > ```bash
> > cat /tmp/test.ini
> > ```
> {: .code-in}
>
> > <code-out-title></code-out-title>
> > The file should look like:
> >
> > ```ini
> > [example]
> > server_name = Dogs!
> > listen = 192.168.0.2
> > apikey = super-secret-api-key-wow!
> > ```
> {: .code-out}
{: .code-2col}
Rendered (try it! resize your browser)
Input: Bashcat /tmp/test.iniOutputThe file should look like:
[example] server_name = Dogs! listen = 192.168.0.2 apikey = super-secret-api-key-wow!
If you leave off the {: .code-2col}, it will render as a single column always.
> <code-in-title>Bash</code-in-title>
> ```bash
> cat /tmp/test.ini
> ```
{: .code-in}
> <code-out-title></code-out-title>
> The file should look like:
>
> ```ini
> [example]
> server_name = Dogs!
> listen = 192.168.0.2
> apikey = super-secret-api-key-wow!
> ```
{: .code-out}
Rendered:
Input: Bashcat /tmp/test.ini
OutputThe file should look like:
[example] server_name = Dogs! listen = 192.168.0.2 apikey = super-secret-api-key-wow!
Quote boxes
> If you don't know where you're going, you might not get there.
{: .quote cite="https://en.m.wikiquote.org/wiki/Yogi_Berra" author="Yogi Berra"}
Rendered:
If you don’t know where you’re going, you might not get there.
The citation and author parameters are both optional. If provided the cite key must be a URL.
If you don’t know where you’re going, you might not get there.
If you don’t know where you’re going, you might not get there.
Additional Features to Improve Learning
Here we cover additional features you can use throughout your tutorials to improve the learning experience.
Tool Links
With the new GTN in Galaxy Webhook, trainees can view training directly within Galaxy. As part of this, we enable those trainees to click on tools, and have those tools directly activated in Galaxy, enabling for a seamless training experience for trainees.
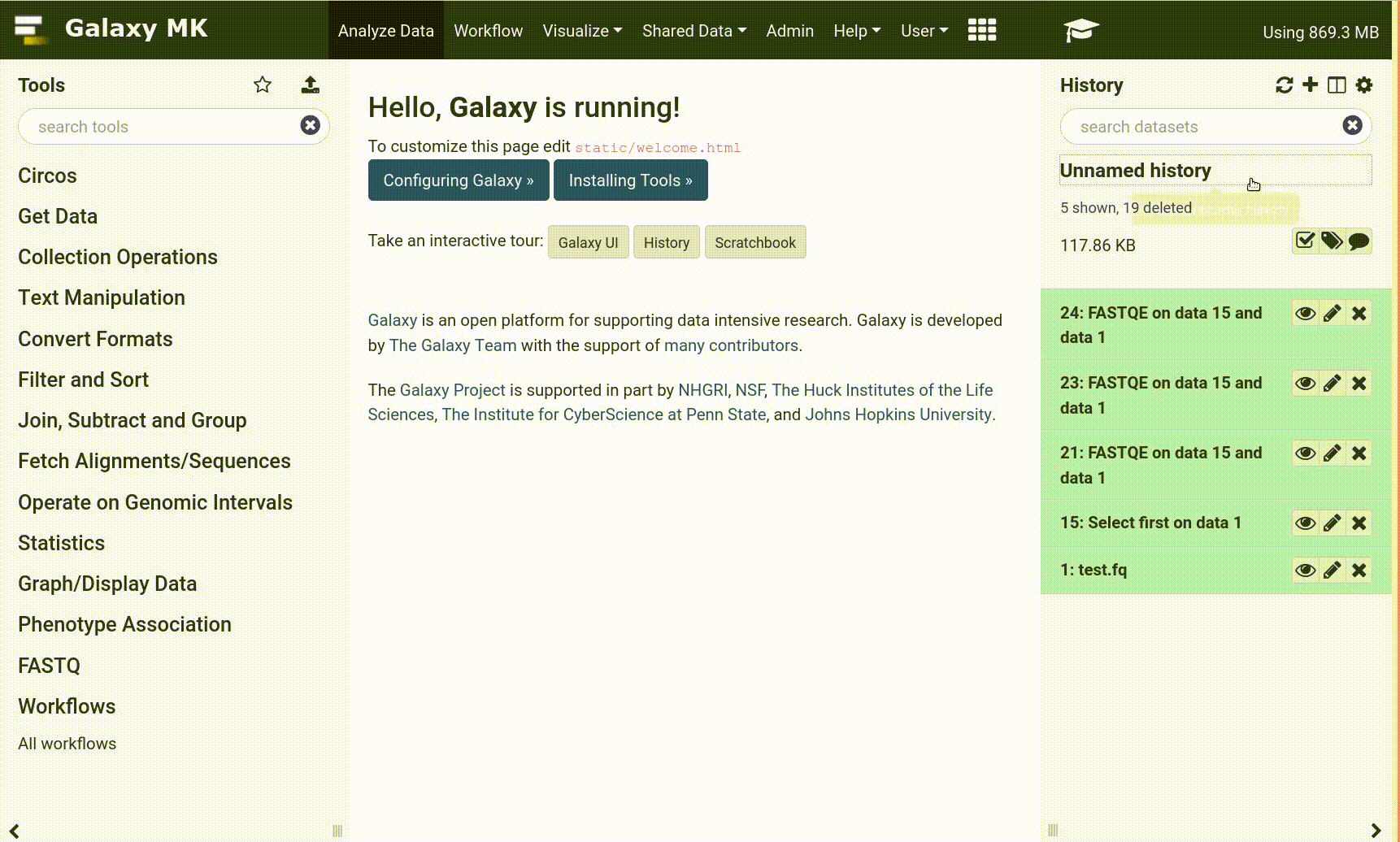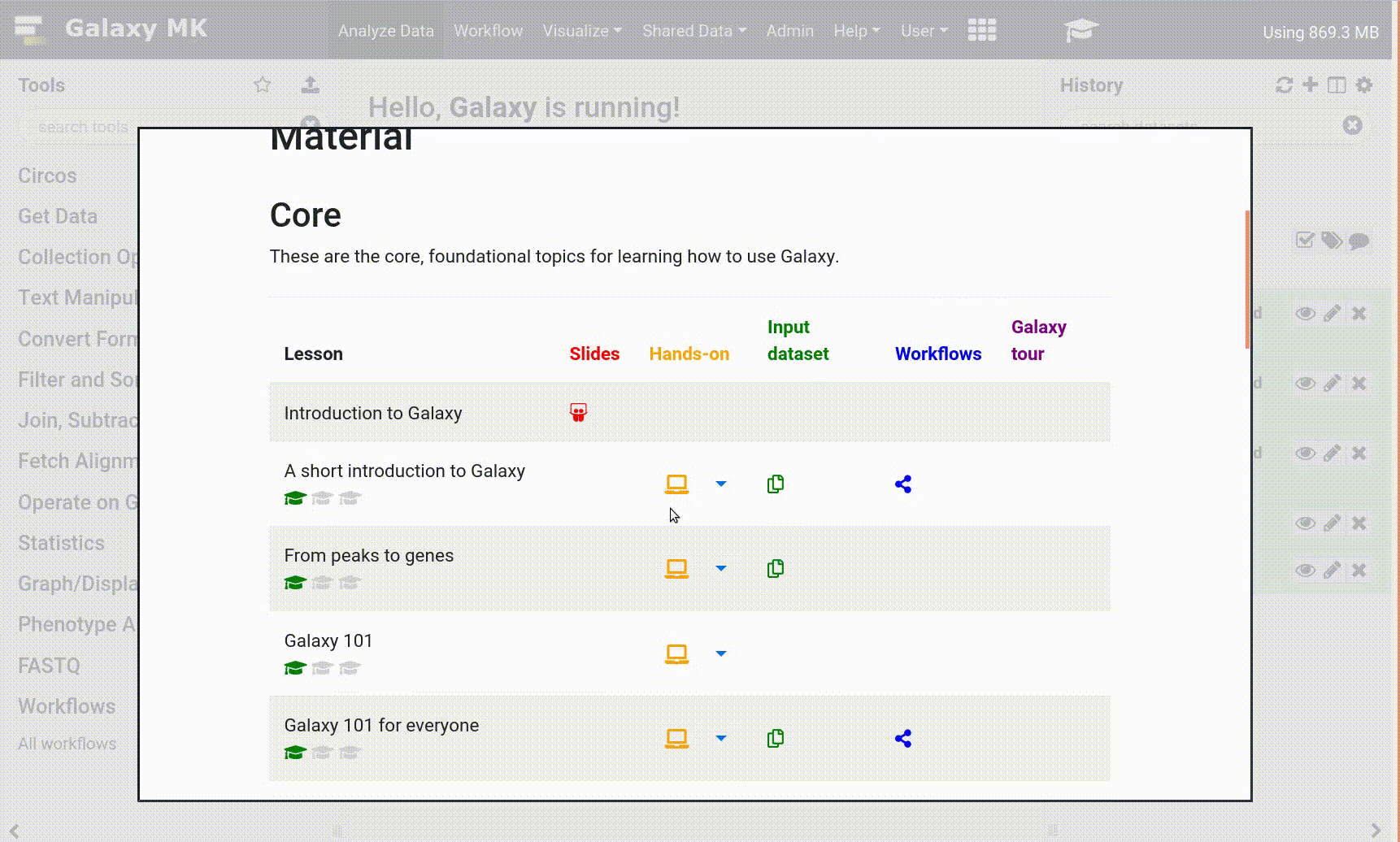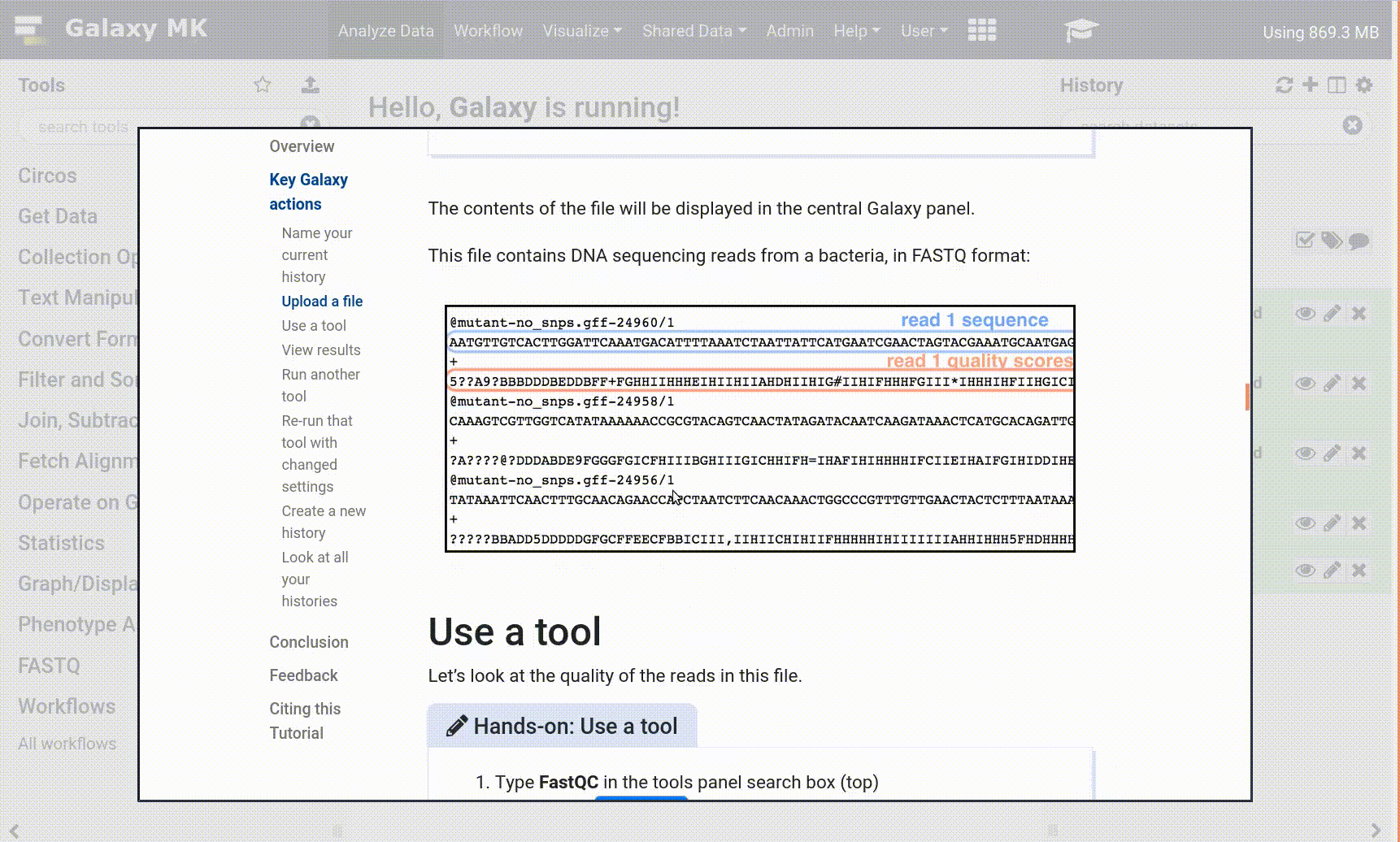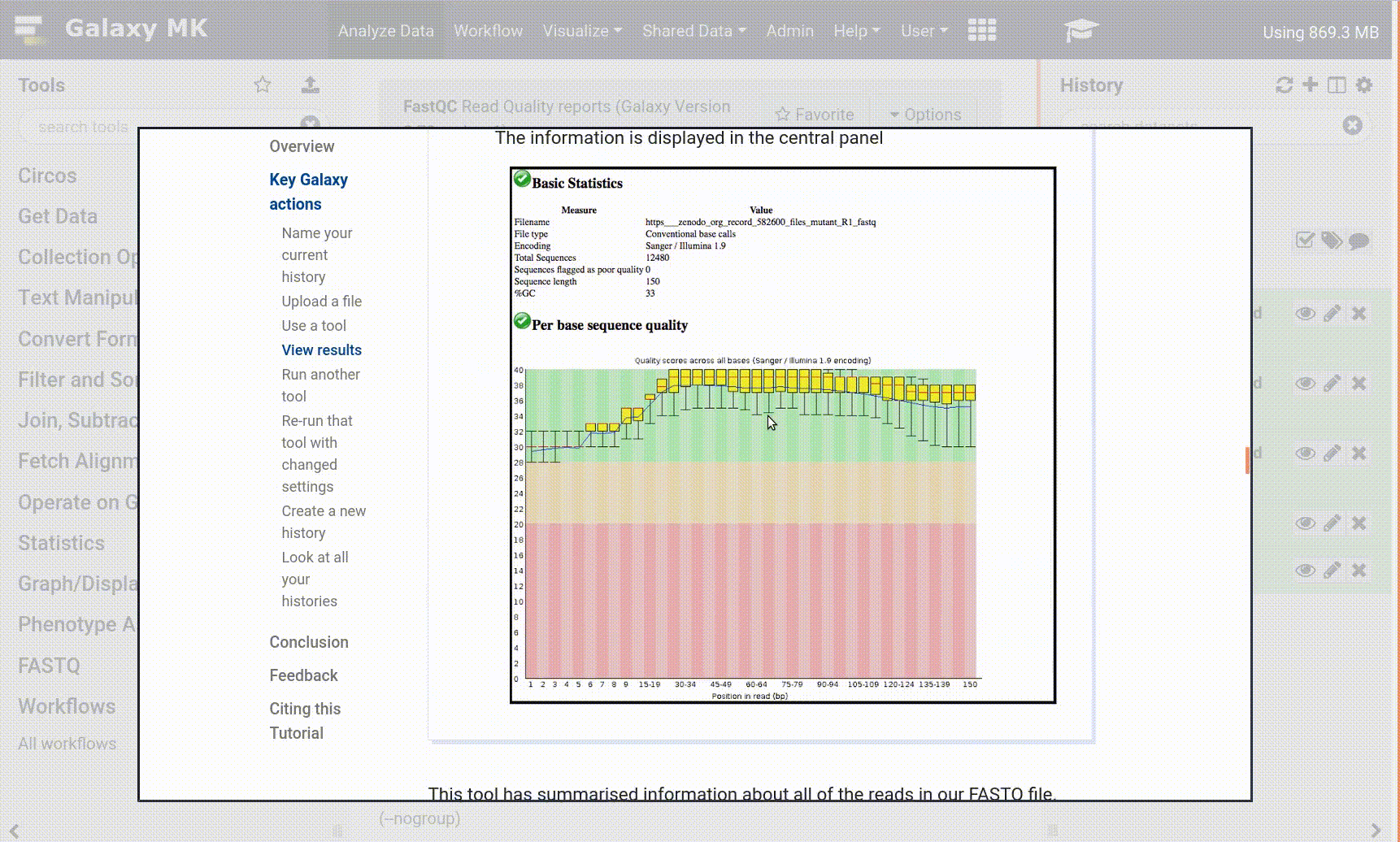 Open image in new tab
Open image in new tabTo enable these in your tutorial you can use the following syntax:
- {% tool MultiQC %}
- {% tool [MultiQC](toolshed.g2.bx.psu.edu/repos/iuc/multiqc/multiqc/1.8+galaxy0) %}
- {% tool [Import some data](upload1) %}
Which will be rendered as:
- MultiQC
- MultiQC ( Galaxy version 1.8+galaxy0)
- Import some data
When viewed through Galaxy, students will see:
Import some data Tool: upload1
How to find these IDs?
The easiest way is to use planemo to generate the training from a workflow. In recent versions of planemo, this is managed automatically.
The alternative is to figure out the ID for the tool you want to use:
- Find your tool in Galaxy, and click to access the tool form.
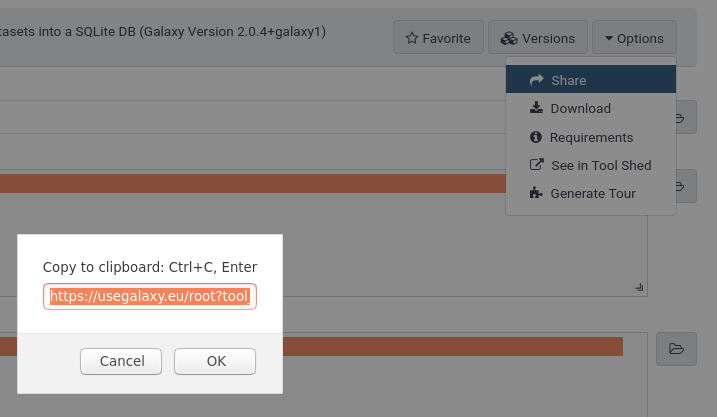
- Click on Options at the top right
- Click on Share
- The URL shown will be something like
https://usegalaxy.eu/root?tool_id=toolshed.g2.bx.psu.edu/repos/galaxyp/mz_to_sqlite/mz_to_sqlite/2.0.4+galaxy1 - Keep only the part after the
=, sotoolshed.g2.bx.psu.edu/repos/galaxyp/mz_to_sqlite/mz_to_sqlite/2.0.4+galaxy1in this example
Workflows
In some tutorials you aren’t as interested in teaching users the individual steps for analysing data, but rather want to focus on some downstream aspects of analysis, or to showcase the best practice workflows that are already available for a user to use! In those cases it can be useful to have a nicer way of inviting the user to execute those steps.
WorkflowHub
You can use a dedicated snippet to invite users to run a WorkflowHub workflow:
{% snippet faqs/galaxy/workflows_run_wfh.md title="mRNA-Seq BY-COVID Pipeline" wfhub_id="685" %}
Rendered:
Hands-on: Importing and Launching a WorkflowHub.eu WorkflowLaunch mRNA-Seq BY-COVID Pipeline (View on WorkflowHub)
Note that it links to a specific workflow, on any Galaxy server. When this tutorial is opened from within the Tutorial Mode, that link will change to one on the current server, removing the intermediate step.
Dockstore
Please note that the dockstore ID should be provided without the # character.
{% snippet faqs/galaxy/workflows_run_ds.md title="My Cool Workflow" dockstore_id="workflow/github.com/jmchilton/galaxy-workflow-dockstore-example-1/mycoolworkflow" %}
Rendered:
Hands-on: Importing and Launching a Dockstore workflowLaunch My Cool Workflow (View on Dockstore)
This snippet has the same behaviour, it will use my.galaxy.training links to make them server independent, but in Tutorial Mode it will open on the current server.
FAQs (snippets)
Many common questions or instructions may be useful to share between different tutorials. For example instructions on how to start a new history or importing data. To make these types of snippets easier to re-use and avoid duplication, they are available in the form of snippets.
The use of snippets is extremely important for asynchronous, remote learning. In this situation as students do not have a teacher immediately on hand, and likely do not have friends or colleagues sitting working with them, they will rely on these boxes to refresh their knowledge and know what to do.
Please ensure you test your learning materials with a learner or colleague not familiar the material, and if possible, (silently) watch them go through your lesson. You’ll easily identify which portions need more explanations and details.
Finding snippets
These are available in folders named faqs, either at the project level, topic level, or tutorial level.
- Project-level FAQs:
faqs/faqs/galaxy/for general Galaxy questionsfaqs/gtn/for questions regarding the GTN website itself
- Topic-level FAQs:
topics/<topic>/faqs/- for questions pertaining to that specific topic
- Tutorial-level FAQs:
topics/<topic>/tutorials/<tutorial>/faqs/- for questions pertaining to that specific tutorial
- if this is present, it is linked to from the tutorial overview box at the top, and from the end of the tutorial
Including FAQs/snippets in your tutorials
To include one of these snippets in your tutorial, you can use the following syntax:
{% snippet faqs/galaxy/histories_create_new.md %}
Which will be rendered as:
Click the new-history icon at the top of the history panel:
The advantage of this approach is that when the Galaxy interface updates, we only have to update the snippet, rather than every tutorial. Please try to use snippets whenever you can!
You could also specify the box type you want the snippet to be rendered in:
{% snippet faqs/galaxy/histories_create_new.md box_type="hands_on" %}
Hands-on: Creating a new historyClick the new-history icon at the top of the history panel:
or without a box altogether:
{% snippet faqs/galaxy/histories_create_new.md box_type="none" %}
Click the new-history icon at the top of the history panel:
Creating new FAQs/snippets
Do you want to include something in your tutorial that you think might be useful in other tutorials as well? Or are you answering a frequently asked question? Consider creating a snippet for it
Each snippet (question) is a separate file, with some metadata, residing in one of the faqs folders:
---
title: How do I run a workflow?
area: workflows # FAQs will be grouped by these areas on the FAQ page
box_type: tip # tip/comment/hands_on/question; optional, if you want the content to be in a box
layout: faq # if you set this the snippet will get its own page and be included in the FAQs page
contributors: [annefou,nomadscientist] # anybody who has contributed to the FAQ over time
---
Here you can write the snippet / answer to the FAQ in Markdown
- Go to `Workflows` on the top menu bar
- Click on ..
- ..
A dictionary/map
Free Textlayout
This must be set to
faqPossible Values:
faqExample(s)
layout: "faq"Free Texttitle
Title of the FAQ
Example(s)
title: How does the GTN implement the 'Ten simple rules for collaborative lesson development'title: How can I give feedback?title: Ways to use GalaxyFree Textbox_type
The type of box that should be used when rendering this FAQ. Use
noneif it should not be in a box, but just plain text.Possible Values:
tiphands_onquestioncommentwarningdetailsnoneExample(s)
box_type: "tip"box_type: "hands_on"box_type: "question"box_type: "comment"box_type: "warning"box_type: "details"box_type: "none"List of Itemscontributors
Who contributed to this FAQ
Example(s)
contributors: - shiltemann - hexylenaFree Text
A text key used for sorting related FAQs together in the visual interface for users.
Example(s)
area: contributorsarea: learnersarea: featuresFree Text
A short, one line description to provide additional context of the FAQ
Example(s)
description: Galaxy may have several reference genomes built-in, but you can also create your own.description: Quickly learn what the identifiers are in any **BAM** dataset that is the result from mappingdescription: Finding and Correcting MetadataList of Items
If a tutorial is renamed to a new location, use this field to list prior locations from which this tutorial was accessible.
Example(s)
redirect_from: - /topics/sequence-analysis/tutorials/de-novo-rad-seq/tutorial
When you are running a remote, asynchronous lesson, you’ll want to be sure you collect all student questions and add them back to your tutorial afterwards, as FAQs. This will help other learners as they progress through the materials, and can give you a very easy URL to point your learners to if they get stuck on a particular task.
FAQ pages
All FAQs will also be collected on their own page, this makes it easy for and teachers to prepare the session, and for participants to quickly find the answers to common questions.
To do this, create a file named index.md inside the faq folder:
---
layout: faq-page
---
If you would like to enforce an order of the faq areas, you can do so:
---
layout: faq-page
area_order: [introduction, learners, instructors, contributors, other]
---
(just make sure you list all existing areas in the folder)
If a tutorial-level FAQ page exists (topics/<topic>/tutorials/<tutorial>/faqs/index.md) it will be automatically linked to from the overview box at the top of the tutorial, and at the end of the tutorial. Have a look at this tutorial to see it in action.
Footnotes
Input: MarkdownFootnotes[^test] can be used to insert more content or explanation as reference material to your content. You can use the same footnote reference multiple time, and the footnote will include backlinks to return to the correct place in the text.[^test]OutputFootnotes1 can be used to insert more content or explanation as reference material to your content. You can use the same footnote reference multiple time, and the footnote will include backlinks to return to the correct place in the text.1
Icons
To use these icons, take the name of the icon, ‘details’ in this example, and write something like this in your tutorial:
{% icon details %}
Some icons have multiple aliases, any may be used, but we’d suggest trying to choose the most semantically appropriate one in case Galaxy later decides to change the icon.
Abbreviations
Oftentimes there are terms you’ll use over and over again where there is an acronym or abbreviation you’ll use consistently. However for learners new to the material, they might need to scroll up to the first definition every time to remember what it meant. It would be annoying as an author to have to re-define it every time, so we’ve implemented some very simple syntax to allow you to create a list of definitions and then use those in the text.
In your tutorial metadata you can add an abbreviations section like:
---
title: My awesome tutorial
...
abbreviations:
API: Application Programming Interface
JSON: JavaScript Object Notation
---
And in your text you can use braces to refer to the term
Input: Markdown
The `/jobs` {API} will return {JSON}. When we call the {API} we'll get back this result {JSON}.OutputThe
/jobsApplication Programming Interface (API) will return JavaScript Object Notation (JSON). When we call the API we’ll get back this result JSON.
A dictionary/map
A dictionary of abbreviations and their expansions.
Example(s)
abbreviations: SQL: Structured Query Language API: Application Programming InterfaceFree Text
The expansion of the abbreviated term.
Choose Your Own Tutorial
Sometimes you’re writing a large tutorial and at one small step there are multiple paths or multiple ways to get the data you want, and you’d like to showcase them all! You could write them all out in order, Option 1…2…etc, however maybe you want it to be a bit more interactive and focus only on one option at a time, so a user doesn’t get distracted by the other options.
Include this markdown where you want your user to choose between the multiple paths:
Input: Markdown{% include _includes/cyoa-choices.html option1="Ananas" option2="Avocados" default="Avocados" text="Here is why some people choose Ananas. Other times you want Avocados as they fit the menu better." %}
Hands-on: Choose Your Own TutorialThis is a "Choose Your Own Tutorial" section, where you can select between multiple paths. Click one of the buttons below to select how you want to follow the tutorial
Here is why some people choose Ananas. Other times you want Avocados as they fit the menu better.
And then they can wrap the relevant sections with a div block with the relevant class. You must set markdown="1" as well to have the inner contents rendered corretly.
NB: If you do not set a default, then on the very first page load, both options will be shown in their entirety. As soon as the user selects one of the options by clicking the relevant button, then the list is filtered. The user’s browser generally will remember which button was selected across navigation and page reloads.
Input: Markdown<div class="Ananas" markdown="1"> - 🍍 are fantastic - hands on! - questions! - solutions! </div> <div class="Avocados" markdown="1"> - 🥑 are amazing - hands on! - questions! - solutions! </div>Output
- 🍍 are fantastic
- hands on!
- questions!
- solutions!
- 🥑 are amazing
- hands on!
- questions!
- solutions!
This can also be used inline: My favourite fruit is an 🍍🥑.
If you wish to have multiple CYOAs in a single tutorial, you are free to do that! However you must:
- Ensure that all options are disjoint, there should not be any shared terms! (I.e if the both CYOAs need to use “STAR”, please find a different way to phrase it, or even use “STAR “, it just needs to be different.)
- Provide a disambiguation term for them, passed as a parameter to all, or all but one, includes.
This disambiguation term will affect the URL parameter, which will become
?gtn-cyoa{term}={value}E.g.:
{% include _includes/cyoa-choices.html option1="Oui" option2="Non" default="Oui" text="Vos données ESTAMP sont prêtes ?" %} {% include _includes/cyoa-choices.html option1="Yes" option2="No" text="Do the thing?" disambiguation="english" %}
URL Parameter
The branch can be selected via URL parameter e.g. for courses, to prevent users selecting the wrong path. Just supply ?gtn-cyoa=Ananas (or your preferred value) on the tutorial URL.
If you are running a remote training, and expect your users to follow a specific path, be certain to include the URL parameter to select the pathway to avoid student confusion. Please note that all tutorials using a CYOA should be tagged which will give you a heads up as a trainer.
Citations
If you would like to cite any articles, books or websites in your tutorial, you can do so by adding a file called tutorial.bib next to your tutorial.md file. In this file you may enter bibtex formatted citations. An example is given below:
@article{bebatut2018community,
title={Community-driven data analysis training for biology},
author={Batut, B{\'e}r{\'e}nice and Hiltemann, Saskia and Bagnacani, Andrea and Baker, Dannon and Bhardwaj, Vivek and Blank, Clemens and Bretaudeau, Anthony and Brillet-Gu{\'e}guen, Loraine and {\v{C}}ech, Martin and Chilton, John and others},
journal={Cell systems},
volume={6},
number={6},
pages={752--758},
year={2018},
publisher={Elsevier},
doi={10.1016/j.cels.2018.05.012}
}
@misc{galaxy-training-materials,
url = {https://training.galaxyproject.org},
note = {Accessed 2019-04-08},
title = {Galaxy Training materials website}
}
@online{Galaxy-P_Metaproteomics,
author = {Galaxy-P Team},
title = {Galaxy-P Metaproteomics instance},
url = {https://proteomics.usegalaxy.eu/},
urldate = {2020-10-13}
}
You can use this in your tutorial as follows:
For more information please look at this great article {% cite bebatut2018community %},
and the corresponding website {% cite galaxy-training-materials %}
Rendered:
For more information please look at this great article Batut et al. 2018, and the corresponding website “Galaxy Training materials website”
A bibliography will automatically be appended to the end of your tutorial (scroll down to the end of this tutorial to see how it looks! or jump there directly)
If you have a DOI for a paper, you can easily obtain the bibtex citation using doi2bib.org.
Automatic Jupyter Notebooks & RMarkdown
If your tutorial is primarily focused on teaching students how to write code (Bash, Python, SQL, etc) you can take advantage of the GTN’s ability to automatically export notebooks from the tutorial content! In this system, you pick a single language for your tutorial, and then all code blocks tagged with that language become runnable. E.g.
Here is some explanation
```python
some_code += f"that students {should execute}"
```
Notebook Schema
A dictionary/map
Example(s)
notebook: language: python pyolite: true notebook: language: python snippet: topics/climate/tutorials/pangeo-notebook/preamble.mdFree Textlanguage
Possible Values:
pythonbashrsqlExample(s)
language: "python"language: "bash"language: "r"language: "sql"List of Items
A list of packages that must be installed before running this tutorial. This value is not currently used, but might be in the future.
Example(s)
packages: - tidyverseBoolean
The GTN has support for JupyterLite and the Pyodide kernel which runs Python in the browser via webassembly/javascript. This comes with some restrictions:
- Python only
- No filesystem access (so no
wgetprep steps)- Little to no cell magic
However, it means we can run a lot of our Python training directly in the GTN! And in the future, hopefully, we will be able to embed individual cells of the notebook directly in the Python training, so the user doesn’t even need to switch pages.
Enabling this field will enable pyolite links for this notebook.
Free Text
If you have an alternative preamble for your notebook that students should know before following (e.g. they must load X datasets in their history), it can be listed here.
This text will be shown in the GTN tutorial, but it will not be included in the notebook, giving you a bit better control over mixing setup content which is relevant for Galaxy, with notebook content that can be easy to run for students.
Example(s)
snippet: topics/climate/tutorials/pangeo-notebook/preamble.md
Currently Supported Languages
| Language | Jupyter | RMarkdown |
|---|---|---|
| Bash | Yes | No |
| SQL | Yes | No |
| R | Yes | Yes |
| Python | Yes | No |
Every cell that you wish to be executable, needs to be annotated with the language like above. Then if a compatible notebook can be produced, it will be.
Restrictions
To use this system, you need to take care of a few things:
- Do not use hands-on boxes for segments that should be executed (code needs to be left aligned!)
- Do not use snippets
- Do not use icons
{% icon galaxy-eye %} - Do not use a terminal or prompt character (that would be included in the execution.)
- Avoid including output when you can, it doesn’t render nicely especially when the cells will become runnable.
And be aware that the output will look a little bit different than the GTN, e.g. solution boxes cannot be hidden by default, so in Jupyter notebook we format the text with a colour of white so it does not appear in the notebook and requires selection to view the answer.
However there are things that are possible! You can still use question/solution boxes, or otherwise nested boxes. Just not includes or snippets
Enabling the system
Add metadata to your tutorial.md header like:
notebook:
language: python
Supported values are python, sql, r, and bash. The notebook will be generated automatically as part of the site build process.
JupyterLite & Pyodide
The GTN has support for JupyterLite and the Pyodide kernel which runs Python in the browser via webassembly/javascript. This comes with some restrictions:
- Python only2
- No filesystem access (so no
wgetprep steps) - Little to no cell magic
However, it means we can run a lot of our Python training directly in the GTN! And in the future, hopefully, we will be able to embed individual cells of the notebook directly in the Python training, so the user doesn’t even need to switch pages.
To enable this feature, set:
notebook:
language: python
pyolite: true
Spanish Translation Project
We have started a trial for translating tutorials into Spanish. Below are instructions on how to add translations of slides and/or hands-on tutorials in Spanish.
- Add a new file with the translated material, next to the English version.
- Add suffix
_ESsuffix- i.e.
tutorial_ES.mdorslides_ES.html
- i.e.
- Add suffix
- Add metadata to the English version (at the top of the file):
tags: - español translations: - es - Add metadata to the translated version of the file:
lang: es translations: - en
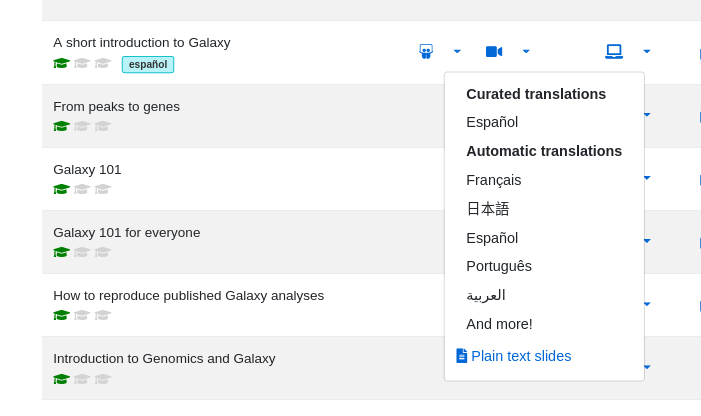
If all worked well, it should look something like this, with a dropdown menu on the slides and/or tutorial showing the presence of a curated tutorial:
Other Languages
Would you like to add a different language to the GTN? Please contact us first (e.g. on Gitter), to discuss a long-term sustainability plan!
Conclusion
If you have created a new tutorial, please also consider writing a GTN news post about it to let people know about it (and make it easy for us to tweet about)!
Footnotes (Rendered)
-
The wikipedia definition of a footnote is: “A note is a string of text placed at the bottom of a page in a book or document or at the end of a chapter, volume or the whole text. The note can provide an author’s comments on the main text or citations of a reference work in support of the text. Footnotes are notes at the foot of the page while endnotes are collected under a separate heading at the end of a chapter, volume, or entire work. Unlike footnotes, endnotes have the advantage of not affecting the layout of the main text, but may cause inconvenience to readers who have to move back and forth between the main text and the endnotes.” ↩ ↩2
-
Not entirely true, other kernels are supported, see their demo repo, but e.g. the SQLite kernel comes with severe restrictions like no downloading databases or connecting to ones online. ↩